Materials Studio Overview Materials Studio is a complete modeling and simulation environment designed to allow researchers in materials science and chemistry to predict and understand the relationships of a material’s atomic and molecular structure with its properties and behavior. Using Materials Studio, researchers in many industries are engineering better performing materials of all types, including pharmaceuticals, catalysts, polymers and composites, metals and alloys, batteries and fuel cells, and more. With Materials Studio you can: Accelerate Innovation: Materials Studio enables materials scientists and research teams to develop new, better performing, and more cost effective materials faster and more efficiently than with test and experimentation alone. Reduce Costs: Materials Studio customers have indicated a reduction of up to 10 times in the number of experiments required to introduce a new material. Improve Efficiency: Automate repetitive or tedious modeling tasks by creating reusable modeling and simulation protocols. Collaborate: Capture & share expert knowledge and methods to make computational science more consistent across organizational and geographic boundaries.
Solve Your Most Difficult Problems: BIOVIA’s staff of expert scientists assures timely support and expertise to help solve even the most challenging problems in materials science. Materials Studio includes a graphical user environment—Materials Studio Visualizer— in which researchers can construct, manipulate and view models of molecules, crystalline materials, surfaces, polymers, and mesoscale structures. Is complemented by a complete set of solution methods including quantum, atomistic (or “classical”), mesoscale, and statistical that enable researchers to evaluate materials at various particle sizes and time scales. It also includes tools for evaluating crystal structure and crystal growth. Materials Studio Integration with BIOVIA Pipeline Pilot Materials Studio is integrated with BIOVIA’s open, scalable, and scientifically aware platform, making it the industry’s most compelling value in materials modeling and simulation. Expert scientists can encapsulate and automate best practices in reusable “protocols” using and its scientific collections, including the.
It can also be integrated with 3rd party or in-house applications to extend the breadth and depth of science supported by Materials Studio. Learn More about. Image Gallery.
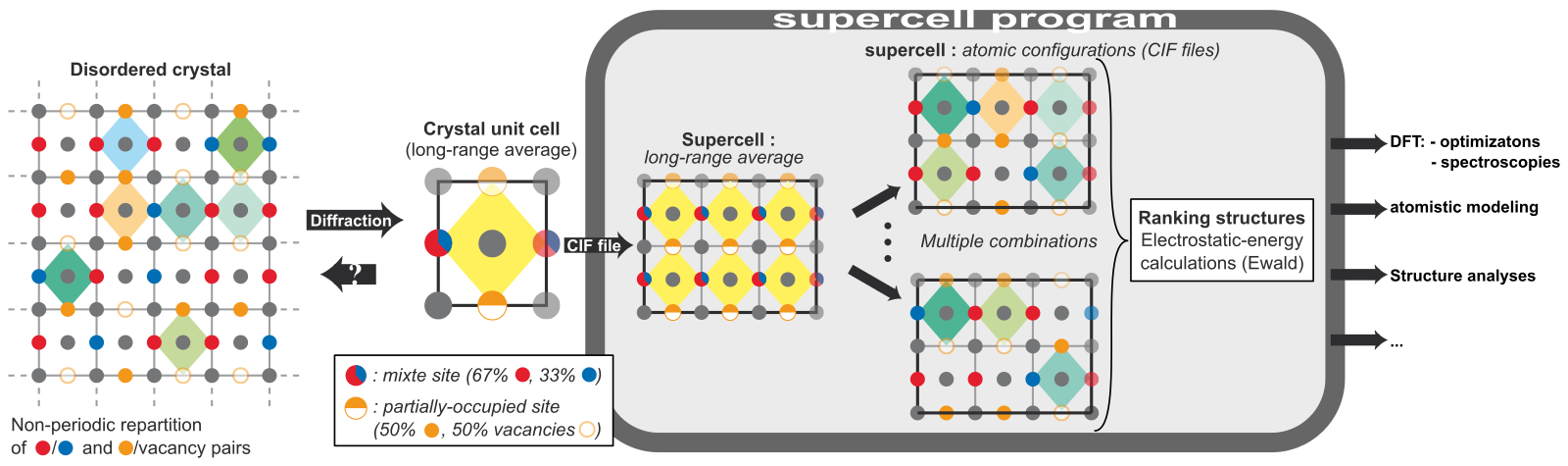
Reaction rate calculated using CASTEP for the dissociation of oxygen on a Pd (111) surface. Examples of different Pt-Ce-O alloy structures generated using the Statistical Disorder tool.
Transmission spectrum for silicon-doped pyrene with aluminium electrodes simulated using electron transport in DMol3. Graphene transport device showing the effect of a nitrogen defect on electron transport. Nanotube transport device showing the effect of a nitrogen defect on electron transport. Angles calculated between best-fit planes for Disperse Red dye give quantitative information on crystal packing.
Estimate solubility of molecules in solvents using free energy of solvation.
Documentation for Castep Utilities Documentation for Castep Utilities pdb2fort (Version 1.0), log2pdb (Version 1.1), log2fort (Version 1.0) and jnl2pdb (Version 1) © Matthew Segall 20/12/94, 11/1/95, 2/2/95, 24/2/95 Note: No guaranty is provided regrading the correctness or suitability of this document or the utilities described herein. Introduction The two applications pdb2fort and log2pdb are for use with CAmbridge Serial Total Energy Package (CASTEP). They enable easy conversion of Protein DataBase (pdb) files into a form suitable for use with CASTEP (pdb2fort) and CASTEP log files into pdb files for easy presentation of results (log2pdb). The pdb filetype is widely used and is recognised by most molecular modeling packages including the public domain application 'Rasmol'. The application jnl2pdb converts.jnl files created by the Cambridge Structure Database (CSD) program Quest into simple pdb files. The CSD is produced by the Cambridge Crystallographic Data Centre located at 12 Union Road, Cambridge, CB2 1EZ. The application log2fort converts a CASTEP log output into a fort.15 CASTEP input file.
There are many potential additions that could be made to these utilities and if you have any ideas please contact the author (address below). Alterantively (preferably) if you want to try to make some changes yourself feel free, but please see the conditions below. If you have any questions regarding CASTEP and its uses or these utilities, please contact the author. Pdb2fort This utility takes a pdb format file and creates a fort.15 file suitable for use with CASTEP.
Further information for inclusion in the param.inc and fort.14 files is also given. To use the utility ensure that an executable is visible in your PATH environment variable and type: pdb2fort at the command line. The atoms read from the pdb file will be displayed along with their positions in Angstroms along orthonormal axis directions. The minimum and maximum x,y and z coordinates are displayed to allow the choice of suitable lattice parameters. The program will then prompt the user to enter the supercell lattice parameters for the x,y and z directions. An orthonormal basis set is assumed and these values should be given in Angstroms in the following format: x,y,z The number of atoms of each type and the species corresponding to that type are then displayed along with the total number of atoms and types and the maximum number of atoms of any one type. This information can be used to set the parameters correctly in the param.inc, fort.14 and fort.10.
A output will be written into which is a complete fort.15 file suitable for use with CASTEP. The only k-point that has been included is the gamma point (0,0,0) with a weight of 1. This is because the primary use for this system is in modeling single molecules which will ideally by dispersionless. This can of course be altered manually by the user. Limitations:. Assumes orthonormal basis for atomic positions. Ignores SCALE statements in pdb files.
Creates fort.15 file with only single k point (0,0,0). Only accepts ATOM statements in pdb file. Cannot decipher HETATOM statements. Log2pdb This creates a pdb file from a CASTEP output. A suitable log file can be created by piping the CASTEP output to a file.
The utility can be used by ensuring the executable is within the PATH environment variable and typing log2pdb where is a valid CASTEP output. The program will print the number of atoms and species found within the log file and the number of iterations covered by the log. The user will then be prompted for the symbol corresponding to each species. This symbol should be one that is contained in the pdb file format specification and should be one or two characters long. The dimensions of the array of supercells to be created will then be requested. This should be given in the form; i,j,k where i is the number of repeated cells in the direction of the first basis vector, j the number for the second and k for the third.
Download minecraft pe appx. For example if only one supercell is required enter 1,1,1 or for an array 2 by 4 by 8 supercells 2,4,8 should be entered. The program will then ask for the iteration for which the positions are required. A pdb file will then be output containing these atomic positions with the filename: N.pdb where N is the number of the iteration selected.
Crtanje korak po korak. The user will then be asked if another iteration is required. The user should respond with the single character 'y' if this is desired, otherwise with the single character 'n'. The pdb file is created with the atomic positions given in Angstroms relative to an orthonormal basis, however the basis used for atomic positions in the CASTEP calculation is not limited to orthonormailty. Limitations:.
The utility will only generate headers and ATOM statements in the pdb file. It will not generate more complex statements such as HETATOM or CONNECT instructions. A maximum of 999 CASTEP iterations can be dealt with at present. Jnl2pdb This converts.jnl files derived from the CSD into pdb files.
Video Editing Software Free Download
The utility can be used by ensuring the executable is within the PATH environment variable and typing jnl2pdb where is a valid CSD.jnl file. Of course the output generated will only be of use if atomic coordinates for the compounds required are contained in the.jnl file.
The program will list the CSD identifier and name of all compounds caompounds found in the file with an index number corresponding to each entry. Simply enter the index number for each compound you wish to create a pdb file for in turn.
Enter 0 to quit the utility. The atoms will appear in the pdb file in the same order as listed in the.jnl file.
Log2fort This application takes a CASTEP log output which is produced by piping the CASTEP standard output to a file and creates a fort.15 file that can be used to create a new CASTEP run. The utility can be used by ensuring the executable is within the PATH environment variable and typing log2fort where is a valid CASTEP output. The program will print the number of atoms and species found within the log file and the number of iterations covered by the log. It will then prompt the user to input the iteration that is required to be output. A fort.15 file corresponding to the atomic positions at the end of that iteration will then be written with the filename N where N is the number of the iteration. The user will then be asked if another iteration is required.
Software Free Download For Windows 7
The user should respond with the single character 'y' if this is desired, otherwise with the single character 'n'. Conditions of Use, Distribution and Alteration This software may be freely copied and distributed provided the following conditions are met:. No charge is made for the source or executable without the author's prior consent in writing. Any alterations or additions to the code are sent to the author at the address given below.
Pc Software Free Download
The distribution is supplied complete with source code and unaltered except where condition (ii) applies, and this documentation file is included. No alteration is made to the version number. The author, or authorised maintainer will release updated versions as and when it is considered necessary. Any alterations made must be added to the list contained within the cource file. Any additional code or alterations sent to the author are considered to have been released into the public domain.
No payment of any kind will be made for additions incorporated into future releases. How to Contact the Author If you want to contact me you can do so at the following address: Matthew Segall Cavendish Laboratory (TCM) Madingley Road Cambridge CB3 0HE E-mail: Please feel free to send any bug reports, comments, criticisms, additions or alterations to me. As you can see there are plenty of enhancements that can be made!